Designing proteins of a certain shape without knowing the specific internal structure ahead of time
Starting with an image of randomly coloured pixels, randomly change some pixels and keep those changes if the image looks more like a banana than it did before. Repeat this process iteratively. Eventually, you can “hallucinate” an image of a banana.
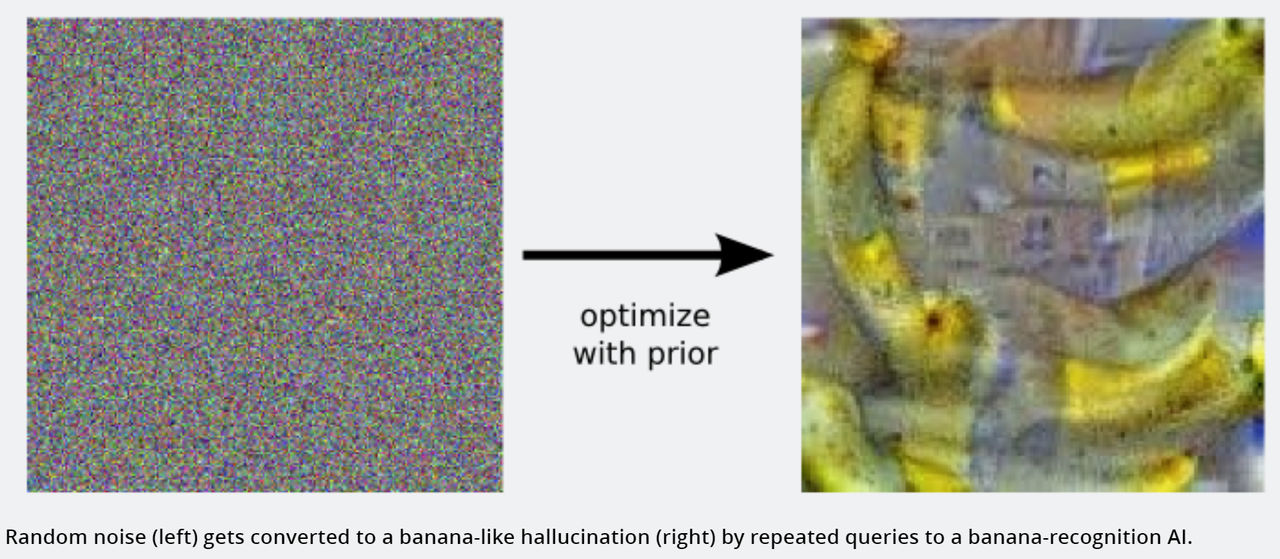

You can also “hallucinate” proteins in much the same manner. Starting with a random sequence of amino acids, randomly change some amino acids and keep those changes only if the protein is more “structured” than it was before. Eventually, you will “hallucinate” a completely novel, ordered protein structure.
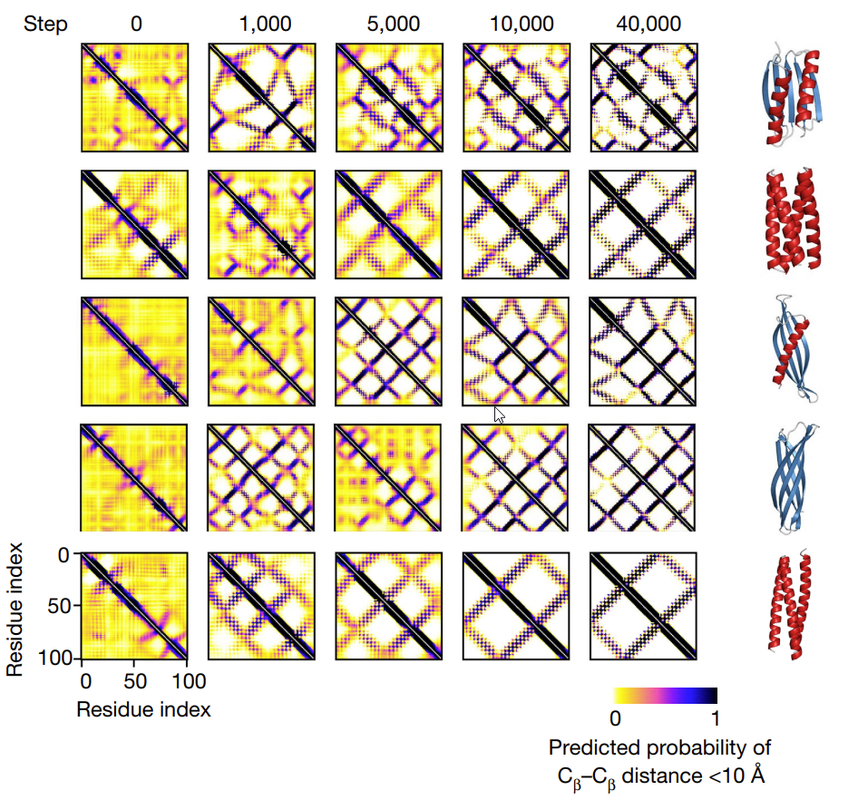
The way you judge if a protein is more “structured” than it was before is to look at the the 2-D contact map. (See image above). Sharp lines on the contact map mean the protein is highly structured.
More recently, Wang et al. 2022 hallucinate proteins scaffolding functional sites. To hallucinate proteins, they minimize the entropy between residue-residue distance and orientation angles, while also ensuring the protein has a desired binding or catalytic motif and the hallucinated structure does not get in the way of it. https://github.com/RosettaCommons/RFDesign
After reading that paper and looking at the GitHub, I had a question. How does one design a protein that forms a certain shape, without knowing the internal structure ahead of time? In principle you can address this using hallucination by defining a loss function that rewards the protein structure fitting in a certain pre-defined space.
To implement this loss function, I represent the pre-defined space as a mesh and then convert it to a cloud of points. The hallucinated protein structure is also represented as a cloud of points. With the Python Open3d library, I can align or “register” the two point clouds and then compute the distance between them. This distance is a measure of similarity between the hallucinated protein structure and the pre-defined space. With this implementation, we can design a protein that forms a certain shape, without knowing the specific internal structure ahead of time.
I have forked the RFDesign GitHub codebase, and I am currently adding in the loss function described above. This is a work in progress. https://github.com/Joseph-Vineland/RFDesign-Shapes
(Thanks to Jue Wang for the correspondence).
To be continued …